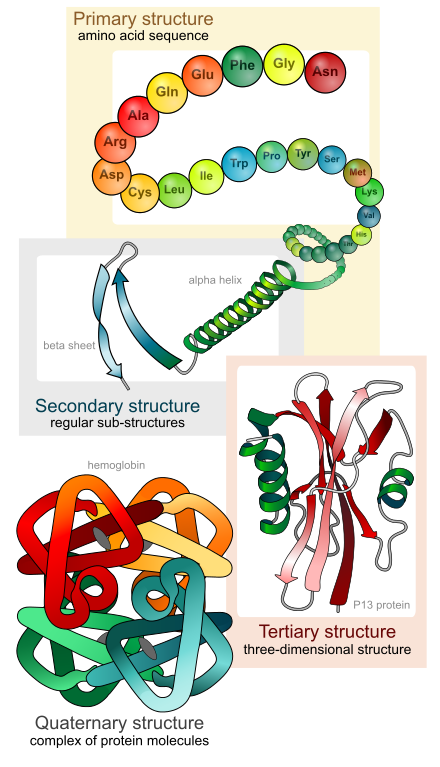
Protein Classifications
Tertiary Structure: the arrangement of secondary structure (sheets, helices, and loops) within a single polypeptide.
Quaternary Structure: arrangement of several polypeptides within a protein complex that performs a specific cellular function.
A dimer: a macromolecular complex formed by two macromolecules. This is an example of a quaternary structure. A homodimer: macromolecules are the same.
Motifs: smallest grouping of secondary structural elements that assemble in a consistent way. These recur in many different proteins with varying sequences. They are conserved.
Examples: β-α-β, two adjacent β-sheets connected by an α-helix. This allows for the creation of a parallel β-sheet. Conserved across proteins.
Domains: Independent folding units within proteins. They can fold on their own and maintain a tertiary structure independent of the rest of the protein. Proteins often have multiple domains, which interact to form functional units. Furthermore, one domain may appear in a variety of different proteins. They are conserved.
Active sites are often in crevasses at the interface between two domains, rich in loops.
Folds: an arrangement of secondary structure elements of a domain or protein: the 3D shape of a particular sequence. Fold factoids:
- Not unique to sequence: different sequences can produce the same fold
- There appear to be a finite number of folds: rate of discovery has plateaued
- Proteins tend to reuse them (folds are conserved)
- To date, roughly 1500 folds have been classified
Domain vs. fold: every domain has a fold, but many folds appear across different proteins.
Similar sequences lead to similar structure, but not always. Often different sequences lead to the same structures too. For example, Hemoglobin and Myoglobin: same basic shape, very different primary sequences.